Multisample Reports#
Multisample reports compare quality metrics across two or more datasets. They enable cross-site comparisons, harmonization assessments, and multi-cohort quality evaluations.
For execution instructions, see the Tutorial.
When to Use Multisample Reports#
Multisample reports are valuable for:
Multi-site studies: Compare data quality across acquisition sites
Longitudinal studies: Track quality changes across collection waves
Harmonization: Assess whether datasets are comparable for pooled analysis
Benchmarking: Compare a new dataset against established references
Report Types#
MEGqc generates two types of multisample reports:
Report |
Focus |
Main Question |
|---|---|---|
QA Multisample |
Signal characteristics |
“How do raw quality profiles compare across datasets?” |
QC Multisample |
Quality scores |
“How do GQI scores and QC components compare across datasets?” |
QA Multisample Report#
Structure#
The QA Multisample report follows the same five-section structure as QA Group, but organized for cross-dataset comparison:
Top tabs: Combined (mag+grad) | MAG | GRAD
└── Section tabs:
├── Summary distributions (merged across datasets)
├── Cohort QA overview (one subtab per dataset)
├── QA metrics across tasks (one subtab per dataset)
├── QA metrics details (shared distributions + dataset-specific heatmaps)
└── Cumulative distributions (pooled ECDFs across all datasets)
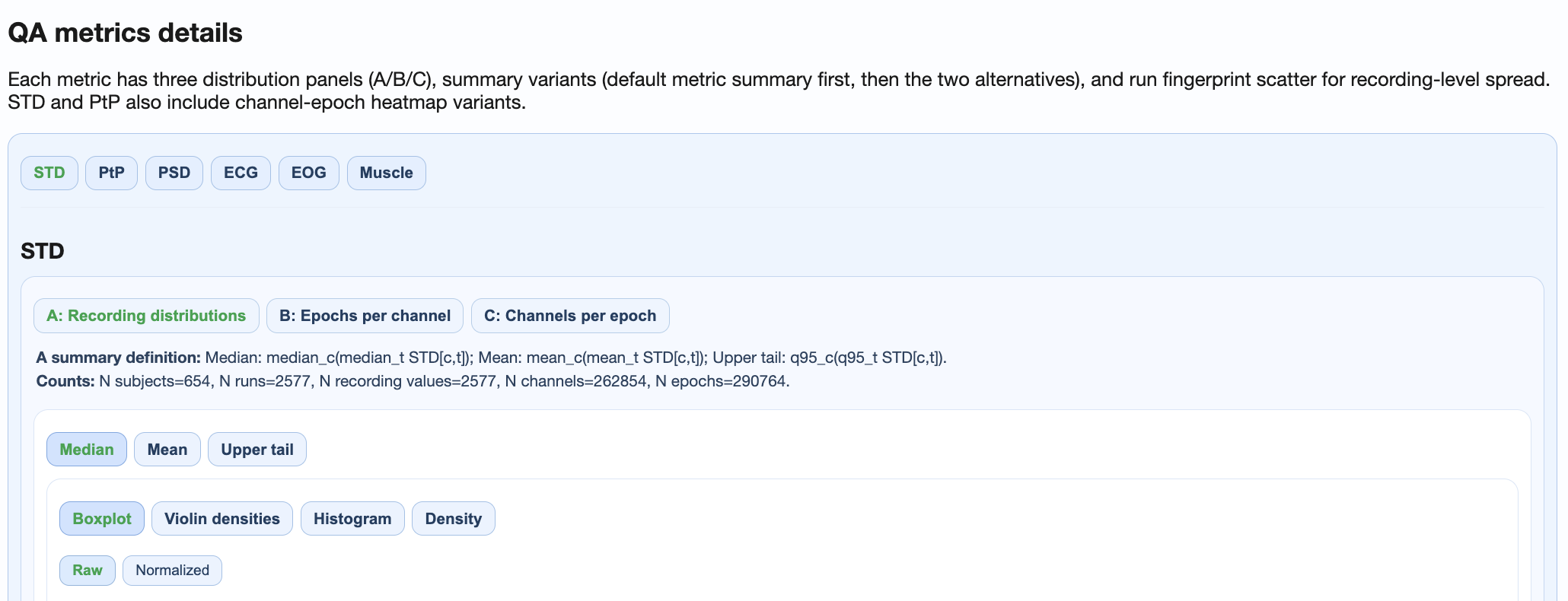
Key Features#
Summary Distributions:
All datasets shown together in violin/box plots
Color-coded by dataset for easy identification
Enables direct visual comparison of distribution shapes
Cohort QA Overview:
One subtab per dataset
Same heatmaps and ranking tables as single-dataset QA Group
Switch between tabs to compare cohort patterns
QA Metrics Details:
Shared plots: Cross-dataset distributions and fingerprints
Dataset-specific subtabs: Heatmaps and topomaps per dataset
Enables both aggregate and detailed comparison
Cumulative Distributions:
All datasets pooled into ECDF plots
Compare distribution tails across datasets
Identify systematic between-dataset differences
Interpretation Tips#
Similar distributions: Datasets are comparable; suitable for pooled analysis
Shifted distributions: Systematic quality differences; investigate causes
Different shapes: Different artifact patterns; may need dataset-specific QC thresholds
QC Multisample Report#
Structure#
The QC Multisample report compares GQI scores and QC components across datasets:
Top tabs: Combined (mag+grad) | MAG | GRAD
└── Metric tabs: GQI | STD | PtP | PSD | ECG | EOG | Muscle
└── Cross-dataset distribution comparisons for each component
Key Features#
GQI Tab:
Side-by-side GQI score distributions per dataset
Penalty breakdown comparisons
Identify which datasets have better/worse overall quality
Component Tabs:
Each QC component compared across datasets
Helps identify which metrics drive between-dataset differences
Support decisions about dataset-specific thresholds
Interpretation Tips#
Similar GQI distributions: Datasets can use the same QC thresholds
Different penalty profiles: Different artifact types dominate each dataset
Consistent component patterns: Similar data quality characteristics
Practical Workflow#
Step 1: Run QA Multisample#
run-megqc-plotting --inputdata /path/ds1 /path/ds2 --qa-multisample
Step 2: Assess comparability#
Check Summary Distributions for overall similarity
Use Cohort QA Overview to identify dataset-specific outliers
Use Cumulative Distributions to compare tails
Step 3: Run QC Multisample#
run-megqc-plotting --inputdata /path/ds1 /path/ds2 --qc-multisample
Step 4: Harmonization decisions#
Compare GQI distributions
Decide on unified vs dataset-specific thresholds
Document quality differences for downstream analysis
Preconditions#
For meaningful multisample comparison:
At least 2 datasets are required
Compatible analysis profiles are recommended (same metrics enabled)
Similar GQI parameterization enables fair score comparison
Comparable tasks/conditions make task-dependent analysis interpretable
Common Use Cases#
Multi-Site Harmonization#
Compare quality profiles across sites to:
Identify site-specific artifacts
Establish site-specific QC thresholds if needed
Document quality differences for publications
Longitudinal Quality Tracking#
Compare data collection waves to:
Detect equipment degradation
Identify protocol drift
Track quality improvements after interventions
Reference Benchmarking#
Compare a new dataset against a well-characterized reference to:
Validate data collection procedures
Establish expected quality ranges
Identify unexpected issues early
